 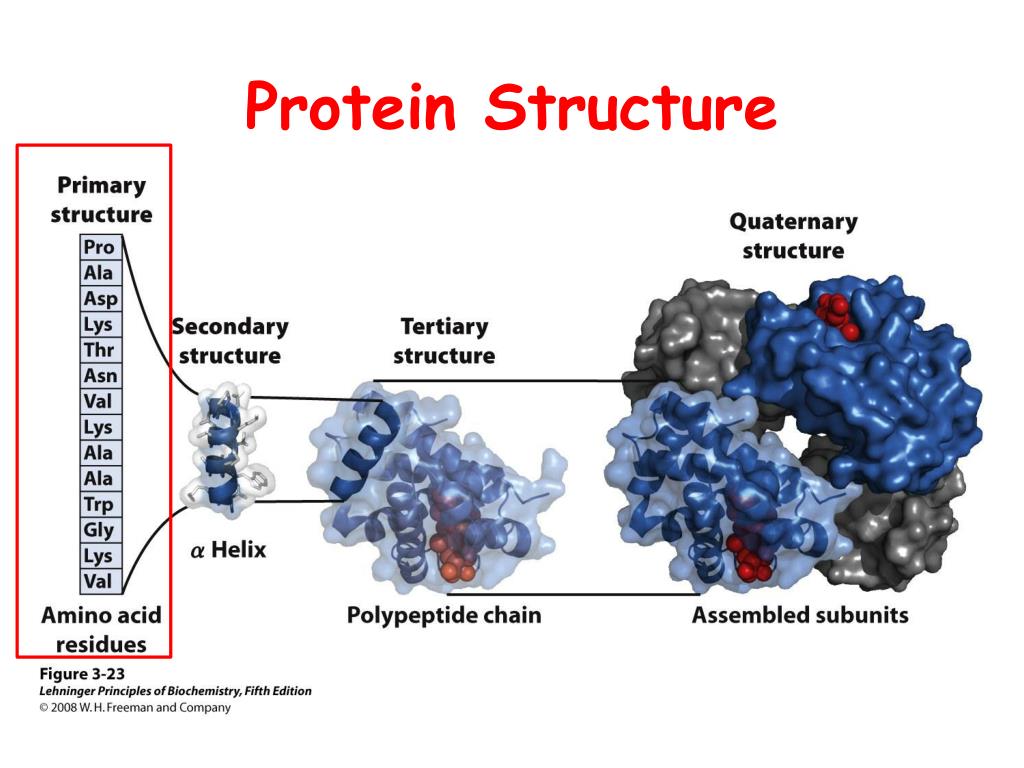
REMARK 3 CROSS-VALIDATION METHOD : THROUGHOUT REMARK 3 COMPLETENESS (WORKING+TEST) (%) : 97.9 REMARK 3 DATA CUTOFF LOW (ABS(F)) : 0.0000 REMARK 3 RESOLUTION RANGE LOW (ANGSTROMS) : 29.55 REMARK 3 RESOLUTION RANGE HIGH (ANGSTROMS) : 2.10 REMARK 3 REFINEMENT TARGET : ENGH & HUBER REMARK 3 : KUNSTLEVE,JIANG,KUSZEWSKI,NILGES, PANNU, REMARK 3 AUTHORS : BRUNGER,ADAMS,CLORE,DELANO,GROS,GROSSE. The result is that parts of the model have not been accounted for, which gives a poor fit to the experimental data. This is due to the high flexibility of parts of the structure, making reliable model building impossible. The R-factor is ok but not great in this case. At this resolution, the electron density for most atoms looks nice and well-separated from its neighbors.īelow is part of a PDB file header showing some of the data like resolution (2.1 Å), resolution range of the data (from lowest, 29.55 Å to highest, 2.10 Å), the number of reflections collected from the crystal during the X-ray experiment (22179) and the R-factor (0,214). In my experience, good quality, well-refined protein structures have a resolution of (or better than) 2.2 Å and an R-factor below 20%. A higher resolution of the X-ray data usually provides a better fit and results in a lower R-factor. The R-factor tells us how good the fit of the structural model to the X-ray data (the electron density) is.  
The R-factor and resolution are probably the two most central parameters for the assessment of the quality of the structure. Apart from the coordinates, the file also contains essential information on the method used to solve the structure, the parameters related to the quality of the X-ray data (like resolution), as well as the symmetry operations for the specific space group of the crystal, the quality of the model geometry (deviations of bond lengths, bond angles and torsion angles from ideal values), secondary structure content, description of missing regions in the structure (a result of weak electron density due to flexibility/disorder in the structure).
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
